Native Isolation of 3×HA‐Tagged Protein Complexes to Characterize Protein‐Protein Interactions - Lim - 2021 - Current Protocols - Wiley Online Library
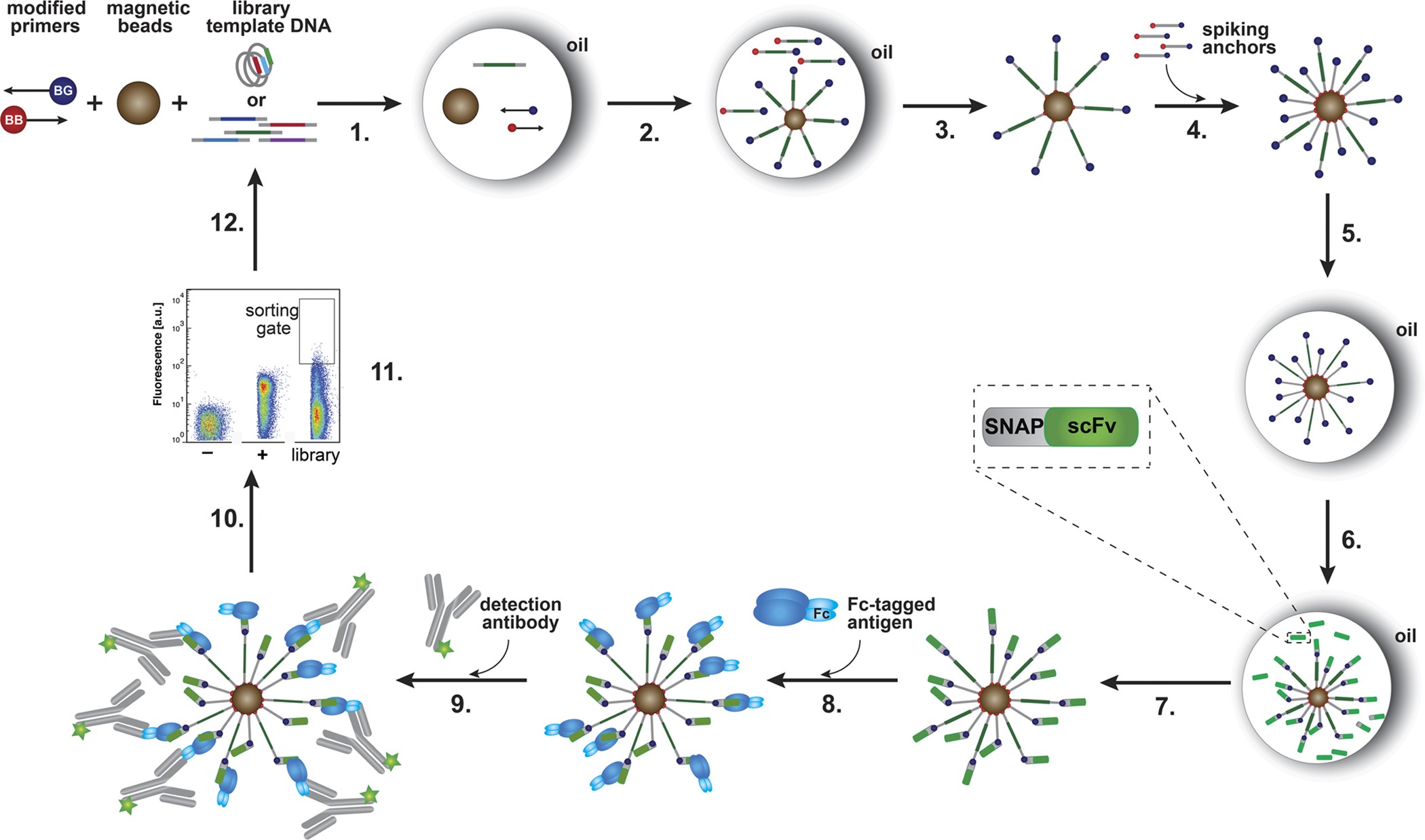
A Shorter Route to Antibody Binders via Quantitative in vitro Bead-Display Screening and Consensus Analysis | Scientific Reports
A proteomics approach to identify targets of the ubiquitin-like molecule Urm1 in Drosophila melanogaster | PLOS ONE
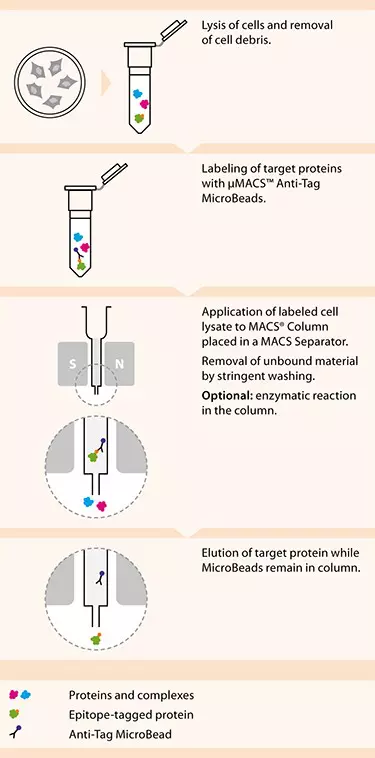
µMACS™ and MultiMACS™ HA Isolation Kits | Epitope-tagged protein isolation and detection | Nucleic acid and protein isolation and analysis | MACS Molecular Analysis | Products and services | USA